 |
| 3D Picture of All Stages of Lactose Operon |
LACTOSE
OPERON:
Many
years ago Francois Jacob and Jacques Monod demonstrated that the bacterial
genes coding for Lactose-metabolizing enzymes are expressed only when lactose
is present .When E.coli cells were grown on a lactose free medium, they did not
produce the enzymes, but they did so within
minutes after being placed on a lactose-enriched medium.Today we know
that a region of E.coli DNA includes a promoter, an operator, and three
adjacent genes associated with lactose metabolism. Any gene(or group of genes),
together with its promoter and operator sequence, is called an operon. The
one we are describing here is the
lactose operon.
 |
| 3D Picture of First Stage of Lactose Operon |
The
regulator gene elsewhere in the DNA codes for a repressor protein that can
inhibit transcription of the Lactose operon. The particular repressor binds
with the operator whenever lactose concentration are low.Being a rather large
molecule, the repressor overlaps the promoter and so blocks RNA polymerase`s access to the
genes.Through the negative control mechanism, Lactose metabolizing enzymes are
not produced when they are not needed.
 |
| 3D Picture of Second Stage of Lactose Operon |
When
lactose is present, however, it binds to and alters the shape of the repressor
protein.In its altersthe shape of the repressor protein.In its altered shape,
the repressor cannot bind to the operator, so RNA polymerase is free to
initiate transcription.
3D animation Showing The Lactose-Operon Stages
When lactose concentrations are high, nearly all the repressor molecules are activated;transcription proceeds rapidly and the Lactose-degrading enzymes are synthesized.
 |
| 3D Picture of Third Stage of Lactose Operon |
 |
| 3D Picture of Fourth Stage of Lactose Operon |
3D animation Showing The Lactose-Operon Stages
When lactose concentrations are high, nearly all the repressor molecules are activated;transcription proceeds rapidly and the Lactose-degrading enzymes are synthesized.
 |
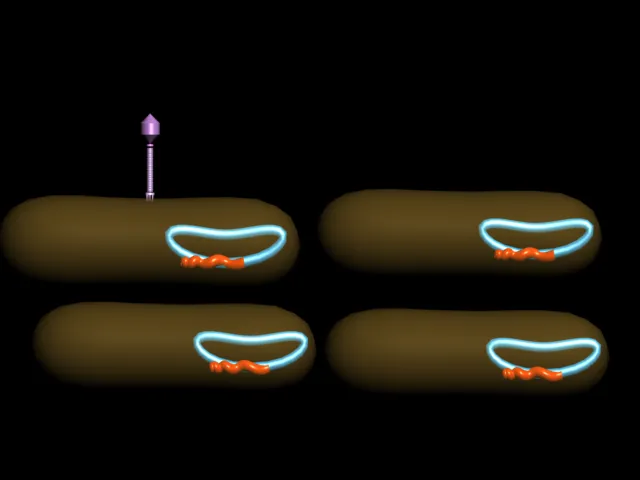
| 3D Picture of A Bacteriophage : is any one of a number of viruses that infect bacteria created by me (Manash Kundu) |
LYTIC AND
LYSOGENIC PATHWAY:
Replication
cycle of Lambda bacteriophage.Depending on environmental factors, infection
proceeds by way of either the lytic pathway or the lysogenic pathway.
 |
| Bacteriophage Lytic Cycle: The lytic cycle is typically considered the main method of viral replication, since it results in the destruction of the infected cell |
In the
lysogenic pathway. The Bacterophage enters a latent state in which the viral
DNA becomes integrated into the host DNA, then remains functionally inactive
during successive DNA replications and cell divisions.
3D Animation Showing Lytic And Lysogenic Cycle of Bacteriophage Virus
Specific environmental agents activate the viral DNA and cause it to leave the host DNA molecule.When it does, the lytic pathway is followed
3D Animation Showing Lytic And Lysogenic Cycle of Bacteriophage Virus
Specific environmental agents activate the viral DNA and cause it to leave the host DNA molecule.When it does, the lytic pathway is followed
For his
studies of inheritance, Morgan relied on the fruit fly Drosophila melanogaster rather
than on the plants and animals that had been commonly used for breeding
experiments.These small flies can be grown in bottles on a little bit of
cornmeal, molasses and agar. A female lays hundreds of eggs in a few days, and
her offspring reach reproductive age in less than two weeks. Thus
Morgan could track hereditary traits through nearly 30 generations of
thousands of flies in a year`s time; Before long, his laboratory was filled
with bottles of busy fruit flies.
 |
| 3D Picture of Normal Drosphila Fly and its Chromosomes created by Me (Manash Kundu) |
 |
| 3D Picture of Mutated Drosophila Fly (White Eyed) and its Chromosomes created by Me (Manash Kundu) |
Drosophila eye color turned out to be a most informative trait. At first all the flies Morgan raised were wild type for eye color; they had brick-red eyes. In 1910, a white-eyed male cropped up in a laboratory bottle. Apparently the variant form arose through a spontaneous mutation in a gene controlling eye color.
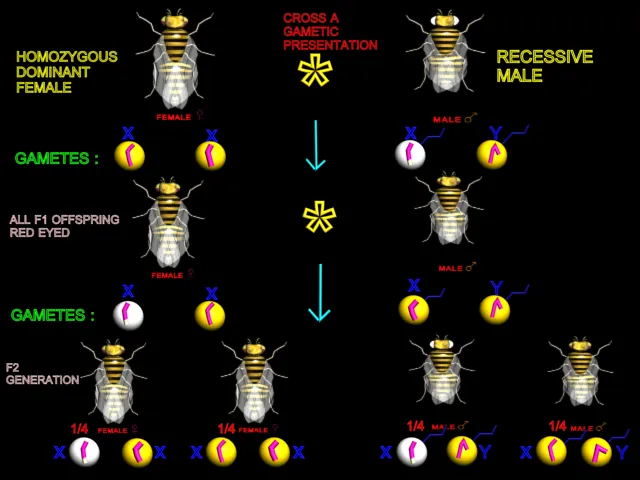 |
| White-eyed males were mated with true-breeding (Homozygous) red-eyed females |
Morgan
established true-breeding strains of white-eyed males and
females. Then he did a series of reciprocal crosses.(In the first of a pair of
crosses, one parent displays the trait in question; in the second cross, the
other parent displays the trait.) White-eyed males were
mated with true-breeding (Homozygous) red-eyed
females. In the reciprocal cross, white
eyed males.The phenotypic outcomes for the paired crosses were not the same.
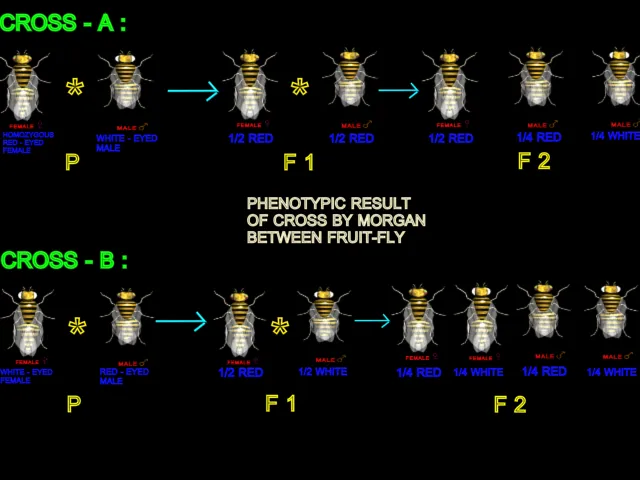 |
| n the reciprocal cross, white eyed males.The phenotypic outcomes for the paired crosses were not the same. |
Clearly
the gene controlling eye color was related to gender, and it probably was
located on one of the sex chromosomes. but which one? Since females
(XX) could be white eyed, the recessive allele would have to be on one
of their X-chromosomes.Now suppose white eyed males
(XY) also carry the recessive allele on their X-chromosome and that
there is no corrosponding eye-colored gene on their Y-chromosome. Those
males would have white eyes beacuse the recessive allele would be the only
eye-color gene they had.
The
results that can be expected when the idea of an X-linked
gene is combined with Mendel`s concept of segregation. By proposing
that a specific gene occurs on the X but not the Y
chromosome,Morgan was able to explain the seemingly odd outcome of his
reciprocal crosses. The results of the experiments matched the predicted
outcomes.
LINKAGE:
During
their Drosophila studies , Morgan and his
co-workers found evidence that many traits are inherited as a group from one
parent or the other.Their studies of an X-linked gene supported the hypothesis
that each gene is located on a specific chromosome, and it seemed likely that
several genes located on the same chromosome might stay physically linked
together during meiosis and ens up in the same gamete.
White
eyes had served as an observable "marker" for tracking the
inheritance of one gene, and now other mutations were used in a similar way.
(By 1915 Morgan`s group has isolated more bthan eighty types of flies with
distinctive mutations.) For example: a wild type fly has straight, flat wings
and gray body, and the two genes for those traits were designated C and B. Some
mutants had curved wings (c) and a black body (b) , and they were mated with
wild type flies in dihybrid crosses. In this type of cross, remember, parents
true-breeding for two traits of interest produce first-generation offspring
that are heterozygous for both traits.All offspring of the CC BB * ccbb cross
were CcBb.
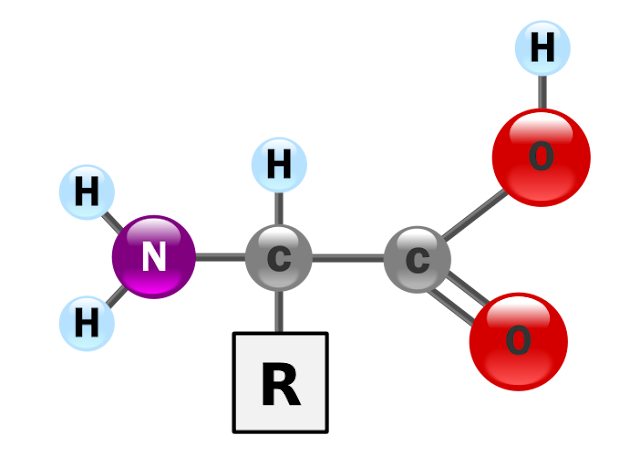
Amino acids
are
molecules containing an amine group, a carboxylic acid group, and a side-chain
that is specific to each amino acid. The key elements of an amino acid are
carbon, hydrogen, oxygen, and nitrogen. They are particularly important in
biochemistry, where the term usually refers to alpha-amino acids.
An
alpha-amino acid has the generic formula H2NCHRCOOH, where R is an organic
substituent;the amino group is attached to the carbon atom immediately adjacent
to the carboxylate group (the α–carbon). Other types of amino acid exist when
the amino group is attached to a different carbon atom; for example, in
gamma-amino acids (such as gamma-amino-butyric acid) the carbon atom to which
the amino group attaches is separated from the carboxylate group by two other
carbon atoms. The various alpha-amino acids differ in which side-chain
(R-group) is attached to their alpha carbon, and can vary in size from just one
hydrogen atom in glycine to a large heterocyclic group in tryptophan.
 |
| Picture of Ten (10) Major Amino Acids |
Amino
acids serve as the building blocks of proteins, which are linear chains of
amino acids. Amino acids can be linked together in varying sequences to form a
vast variety of proteins. Twenty amino acids are naturally incorporated into
polypeptides and are called proteinogenic or standard amino acids. These 20 are
encoded by the universal genetic code. Nine standard amino acids are called
"essential" for humans because they cannot be created from other
compounds by the human body, and so must be taken in as food.
Amino
acids are important in nutrition and are commonly used in nutrition
supplements, fertilizers, food technology and industry. In industry,
applications include the production of biodegradable plastics, drugs, and
chiral catalysts.
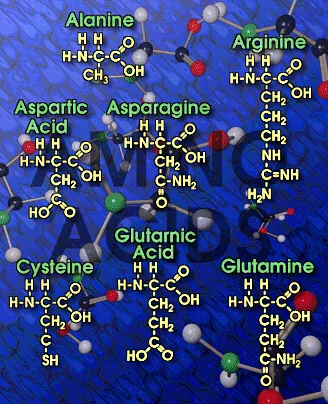 |
| Picture of Major Amino Acids |
Standard
amino acids
Amino
acids are the structural units that make up proteins. They join together to
form short polymer chains called peptides or longer chains called either
polypeptides or proteins. These polymers are linear and unbranched, with each
amino acid within the chain attached to two neighboring amino acids. The
process of making proteins is called translation and involves the step-by-step
addition of amino acids to a growing protein chain by a ribozyme that is called
a ribosome. The order in which the amino acids are added is read through the
genetic code from an mRNA template, which is a RNA copy of one of the
organism's genes.
Twenty-two
amino acids are naturally incorporated into polypeptides and are called
proteinogenic or natural amino acids.Of these, 20 are encoded by the universal
genetic code. The remaining 2, selenocysteine and pyrrolysine, are incorporated
into proteins by unique synthetic mechanisms. Selenocysteine is incorporated
when the mRNA being translated includes a SECIS element, which causes the UGA
codon to encode selenocysteine instead of a stop codon.Pyrrolysine is used by
some methanogenic archaea in enzymes that they use to produce methane. It is
coded for with the codon UAG, which is normally a stop codon in other organisms
This UAG codon is followed by a PYLIS downstream sequence.
Non-standard amino acids
Aside
from the 22 standard amino acids, there are many other amino acids that are
called non-proteinogenic or non-standard. Those either are not found in
proteins (for example carnitine, GABA), or are not produced directly and in
isolation by standard cellular machinery (for example, hydroxyproline and
selenomethionine).
 |
| Amino Acids Page in My Bio-World Software |
THE GENETIC
CODE
AMINO
ACIDS RNA
CODON
ALANINE
GCA GCC GCG GCU
ARGININE AGA AGG
CGA CGC CGG CGU
ASPARAGINE AAC
AAU
ASPARTIC
ACID GAC
GAU
CYSTEINE UGC
UGU
GLUTAMIC
ACID GAA GAG
GLUTAMINE CAA
CAG
GLYCINE
GGA GGC GGG GGU
HISTIDINE CAC
CAU
ISOLEUCINE AUA AUC
AUU
LEUCINE
UUA UUG CUA CUC CUG CUU
LYSINE
AAA AAG
METHIONINE AUG
PHENYLALANINE UUC
UUU
PROLINE CCA CCC
CCG CCU
SERINE
AGC AGU UCA UCC UCG UCU
THREONINE ACA ACC
ACG ACU
TRYPTOPHAN UGG
TYROSINE UAC
UAU
VALINE
GUA GUC GUG GUU
STOP
CODONS UAA UAG UGA
Catabolism:
Degradation
of an amino acid often involves deamination by moving its amino group to
alpha-ketoglutarate, forming glutamate. This process involves transaminases,
often the same as those used in amination during synthesis. In many
vertebrates, the amino group is then removed through the urea cycle and is
excreted in the form of urea. However, amino acid degradation can produce uric
acid or ammonia instead. For example, serine dehydratase converts serine to
pyruvate and ammonia. After removal of one or more amino groups, the
remainder of the molecule can sometimes be used to synthesize new amino acids,
or it can be used for energy by entering glycolysis or the citric acid cycle,
 |
| Picture of All Major Amino Acids usefull during Kreb`s Cycle : Amino Acids are written in yellow over red background |
3D Animation Showing Major Amino-Acids






























 Online Movies
Online Movies
No comments:
Post a Comment